OmopStudyBuilder
Reproducible Multi-Site OMOP Network Studies
OmopStudyBuilder
Background
- Multi-site OMOP studies face long-term reproducibility challenges.
renvhelps short-term collaboration, but long-term reproducibility can still break.- Common failure points include:
- Different R versions used at restore time.
- Operating system and software differences across partner sites.
- Hidden system requirements such as Java, curl, and other shared libraries.
- Containers improve portability, but not every data partner can adopt Docker because of local IT and security restrictions.
What is renv?
Records exact R package versions so any partner can restore the same study environment.
- Records the exact version of every R package used by the study.
- A partner runs
renv::restore()and reinstalls the same package set with no version drift. - The environment is described in a single
renv.lockfile that travels with the study code.
What is Docker?
Docker is a shipping container for the full software environment.
- Packages study code, R version, and system libraries into a single image.
- A partner runs the image and gets the same OS-level runtime, R version, and package stack.
- The image can also expose RStudio Server, so partners still work in a familiar interface.
Docker must be installed on the machine to use this option.
Where This Sits
Arachne
- UI-driven platform for network studies.
- Requires setup, hosting, and maintenance.
- Limited flexibility for study code.
Ulysses
- Templates study execution workflows.
- Closer to a Strategus-style execution model.
- Limited flexibility (more flexible than Arachne).
OmopStudyBuilder
- Lower-level, code-first approach.
- Standardises the project itself.
- Supports both
renvand optional containerisation.
What OmopStudyBuilder Does
- An opinionated R package that standardises how studies are:
- Created with a consistent project structure and templates.
- Validate the renv file (packages from CRAN, no unnecessary dependencies).
- Distributed as a shareable folder with optional Docker image publishing.
- Ships templates for:
- Study code.
- Diagnostics code.
- Shiny applications.
Workflow
The Workflow
Create study -> Write study code -> Record dependencies -> Review -> Build image -> Run -> Share
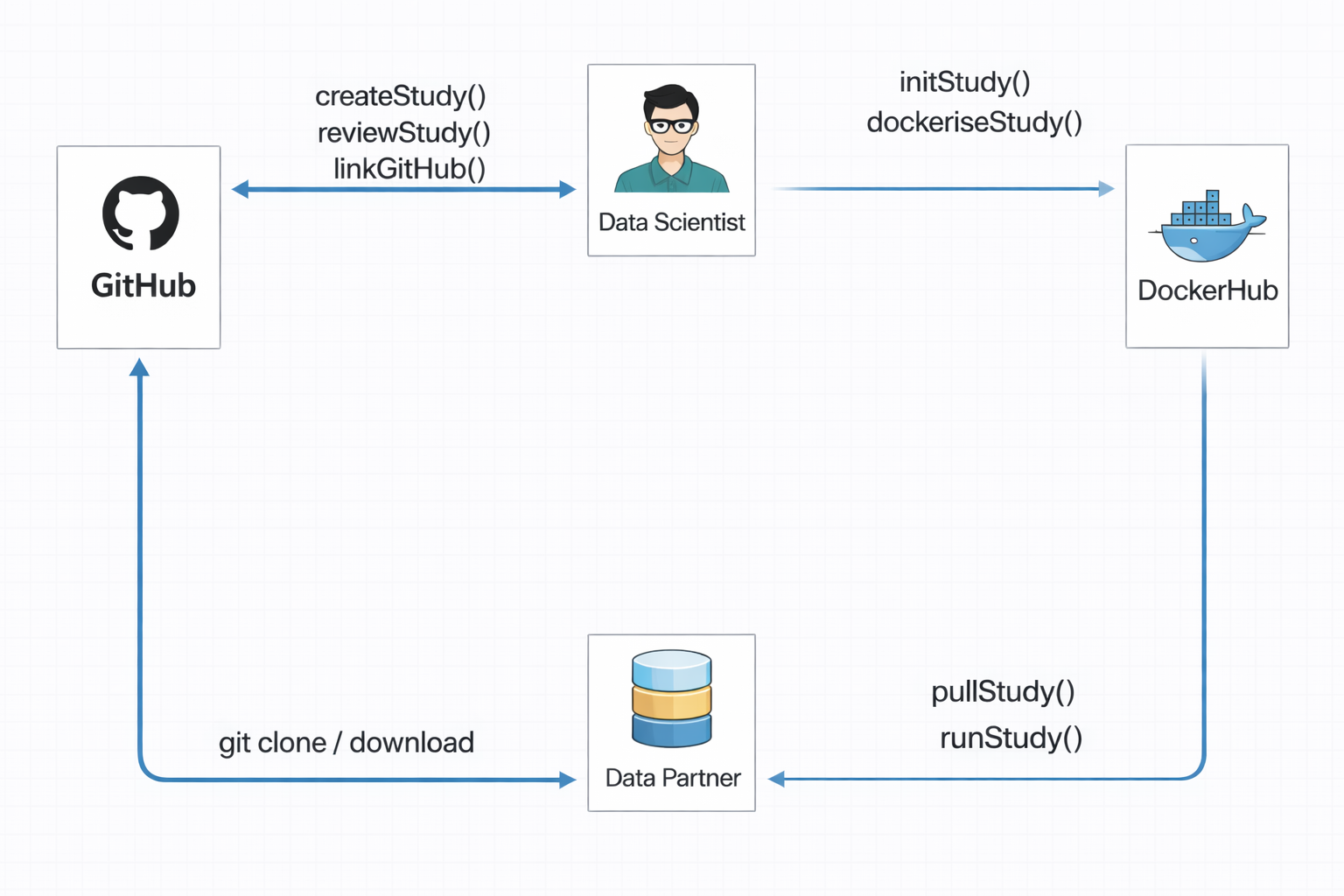
- Data partners can choose their execution mode.
- Option A: run from source with
renv.lock. - Option B: run the Docker image for stronger long-term reproducibility.
- Docker commands are wrapped in R functions, so users do not need to work directly with the CLI.
Why Two Execution Modes?
- Some partners cannot install or run Docker.
- Some studies need a portable runtime across sites for long-term reuse.
- OmopStudyBuilder keeps both paths available in the same study package.
Approach
Create Study
- Creates the initial directory structure for an OMOP CDM network study.
- Gives teams a standard starting point instead of custom folders for each project.
Write Study code
The data scientist then writes the study-specific analysis code inside the generated project structure.

Review
- Reviews dependencies.
- Summarises dependencies captured in
renv.lock. - Helps identify missing requirements before a study is distributed.
Docker
- Builds a Docker image for the study.
- Pushes that image to Docker Hub or another registry for partner reuse.
Typical Workflow
- The package supports the study lifecycle from setup through execution and sharing.
Automated Execution
- Runs the R Docker image for automated execution.
- Expects a
.envfile in the project root for runtime configuration.
Interactive Execution
- Runs RStudio Server for interactive study execution.
- Opens in the browser with the required packages already installed.
What This Improves
- Site differences are reduced when partners use the same image.
- The runtime stays aligned across:
- R version.
- Operating system and system dependencies.
- Installed R packages.
- OmopStudyBuilder smooths over practical rough edges such as:
- Docker availability checks.
- Automatic port selection.
- Clearer execution errors.
OmopStudyBuilder
OmopStudyBuilder: reproducible OMOP network studies